Office of Research & Development |
 |


In VA’s Million Veteran Program, researchers are mapping the human genome by using genotyping—a process that spells out several hundred thousand data points, one-by-one. (Photo for illustrative purposes only © Getty Images/alanphillips)
May 19, 2022
By Mike Richman
VA Research Communications
"The importance of this cannot be overstated. Using TOPMed's imputation panel vastly enriches MVP's genetic database."
Since 2011, more than 875,000 Veterans have donated their DNA to the VA Million Veteran Program (MVP). In the program, VA researchers are sequencing Veterans' genomes—a complete set of genetic material—to better understand how genes, lifestyles, and military exposures can affect a person’s health and risk for illness.
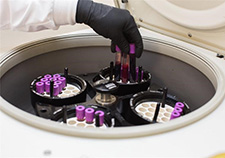
Centrifugation helps scientists extract DNA from a Veteran's blood sample and prepare the DNA for genetic analysis, either through genotyping or whole genome sequencing.
Researchers map the genomes using genotyping—a process that spells out several hundred thousand data points, one-by-one. To fill in any missing data, scientists look to genomes where all data points have been fully sequenced. These genomes are used as a reference to determine the missing letters.
MVP has created whole genome sequences for over 140,000 Veterans. However, it takes time to process this large amount of data and make it available for research. It takes even more time to develop a calculation tool—called an imputation panel—to determine the missing data points that remain when genotyping is complete.

VA investigator brings diversity into autoimmune disease research

Veterans help find new cancer treatments

Million Veteran Program director speaks at international forum
In efforts to find a more accurate and cost-effective method for predicting missing genomic data points, VA investigators found a potential interim solution just across town in Bethesda, Maryland: a genomic research program at the National Institutes of Health (NIH).
As part of that solution, MVP has been collaborating with NIH to use a reference panel for imputation.
The panel is run by TOPMed—the Trans-Omics for Precision Medicine Program. The program, funded by NIH’s National Heart, Lung, and Blood Institute (NHLBI), aims to generate scientific resources that will improve the understanding of heart, lung, blood, and sleep disorders and advance precision medicine, an approach in health care that takes into account a person’s gene variants and his or her environment and lifestyle to find the right treatment.
Since 2014, TOPMed has collected the entire genome sequence of more than 100,000 people. Scientists created a new reference genome and imputation panel based off these whole-genome sequences.
This first-of-its kind imputation panel offers far more statistical accuracy than any previous imputation panels. In that way, it will help scientists develop better treatments specific to a person’s genes and environment.
MVP wanted to find a way to use this tool to improve its genetic database, now one of the largest in the world. The database supports the work of more than 500 researchers across VA. With better genetic data, the researchers can uncover new genetic markers associated with disease in Veterans that could someday revolutionize their health care. As such, MVP leaders set out to strike up a partnership with their colleagues in Bethesda.
In 2018, TOPMed and VA began discussions with the goal of using this new imputation panel to re-do genetic analysis on DNA shared by hundreds of thousands of Veterans in MVP.
“The importance of this cannot be overstated,” says Dr. Phil Tsao, MVP’s co-principal investigator in data generation and access. “Using TOPMed’s imputation panel vastly enriches MVP’s genetic database. Over 500 researchers use MVP data to find new genetic markers related to health and illness in Veterans. With better data, they can make better findings. With better findings, we can offer Veterans better care with more laser-focused accuracy.”
The TOPMed reference genome for this imputation panel was selected with racial and ethnic diversity in mind. This means VA can make much more accurate sequencing predictions for people who come from non-European ancestry. This is critical for MVP, since approximately 30% of the more than 875,000 Veterans in MVP are from non-European descent.
“To be able to more accurately assemble their genome means the research we do on non-European Veteran populations will be much more thorough,” says Dr. Sumitra Muralidhar, the director of MVP. “That means we can discover new, never-before-known genetic markers connected with health and disease in Veteran populations of non-European descent.”
In 2020, VA acquired the TOPMed imputation panel from NIH, and researchers began methodically re-analyzing and updating more than 650,000 genomes from Veterans who provided blood samples when they joined MVP. In December 2021, the update was complete and the new, upgraded data on hundreds of thousands of Veterans became available to research.
“Due to our collaboration with TOPMed, we have better genetic data now, which will hopefully lead to findings of new genetic markers for diseases, specifically in diverse Veterans,” Muralidhar says. “The end result will hopefully be new discoveries and better treatments, as well as a better understanding of the biology of diseases. That way, we can offer more precise treatments for diseases and other health conditions.”
Now that MVP’s genetic data is updated with TOPMed’s reference panel, any findings from MVP data can be compared and validated with findings from other research based on genomes that were also imputed using TOPMed’s reference panel.
“Use of the TOPMed reference panel in MVP, which is one of the world’s largest genomic biobanks, allows for more collaboration between NIH and VA scientists to cross-validate each other’s analysis results,” says Dr. Saiju Pyarajan, director of the Center for Data and Computational Sciences at the VA Boston Healthcare System. The more people around the world who have been compared to the same reference sample, the more we’re able to drastically increase the confidence in the conclusions or results from these genomic analyses. This is wonderful for the future of precision medicine.”
VA Research Currents archives || Sign up for VA Research updates